Biological Cells Segmentation
The task of accurate cell segmentation is essential for cellular biology and single-cell analysis, as well as for studying biological processes as a whole. In biomedical image processing, this includes reconstruction of microscopy images, foreground segmentation, cell detection, cellular compartments and organelles segmentation. Despite the tremendous progress in microscopy cell imaging and numerous segmentation methods, accurate segmentation and reliable characterization of cells remain challenging tasks that usually require problem-specific tailoring of algorithms.
The aim of our project was to develop an automatic solution for specific biological cell segmentation. Namely, it was necessary to develop a robust and precise algorithm for the automatic structural segmentation of cells containing β-catenin, a multifunctional protein that plays a crucial role in the regulation and coordination of cell-cell adhesion and gene transcription. β-catenin is expressed in different cell types and tissues and, depending on its subcellular distribution and expression levels, its imbalance often results in disease and deregulated cell growth connected to cancer and metastasis.
There are some issues with the segmentation of biological cells. It is often difficult to assess and determine cell boundaries visually. Some cells’ boundaries are partially invisible, some cells are overlapped. An example of a source image you can see below in Fig.1:

Fig. 1. Source cell image
In order to develop a high-precision algorithm of cell segmentation, we applied the following scheme:

First and foremost, we improve the visual quality of the source cell image using the Adaptive histogram equalization algorithm. Also, we use some additional information to supplement some missing and invisible cell boundaries on a source image. In addition, very simple image filtration was applied to remove unnecessary cell candidates.
The main steps of image filtration are next:
- Gaussian smoothing to remove some noise;
- CLAHE to highlight cell boundaries;
- A series of morphology operations, such as opening, closing, fill holes, and others;
- Cells clustering and filtration using geometric features.
You can see noticeable changes in cell processing below.

Fig. 2. Result of cells improving and filtration
To improve the segmentation algorithm and localize items more accurately, DAPI (fluorescent probe for staining of DNA) images were used.

Fig. 3. DAPI fluorescent image
Watershed segmentation was applied to the DAPI images to obtain markers (labels) for the characterization of cells with nuclear accumulation of β-catenin.
Finally, DAPI markers were used for final cell segmentation with an improved watershed algorithm.
Below you can see a simple animated demo illustrating how biological cell segmentation works.
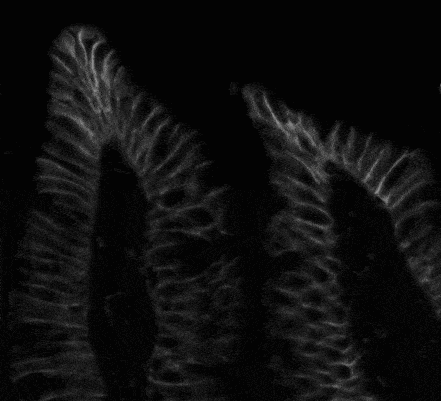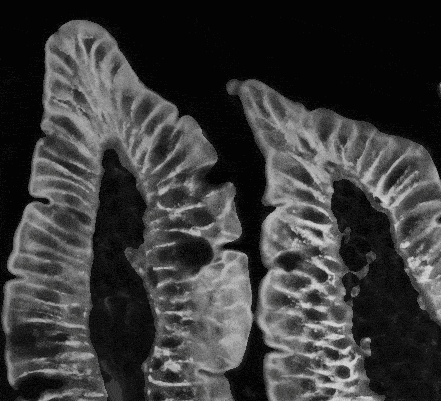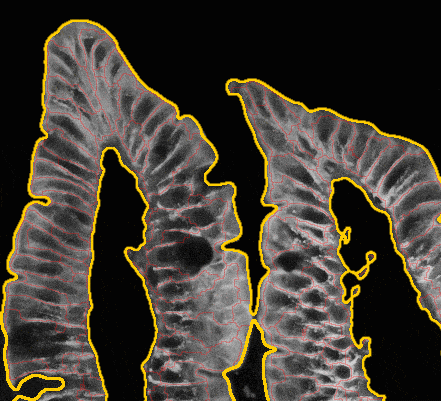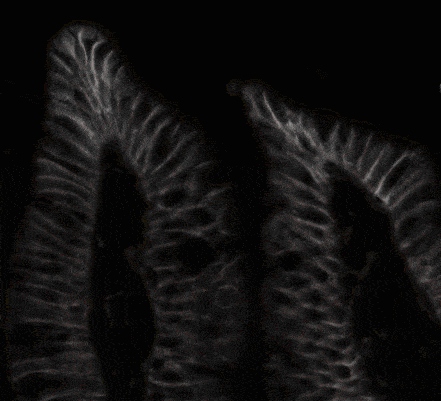
Fig. 4. Сell segmentation demo
In conclusion, being an important technique for studying biological processes, microscopy image processing remains a bottleneck in the analysis of living cells, as it implies the subjective and time-consuming intervention of human experts. Meanwhile, computer vision approaches for segmenting and analyzing the cells can address some of these challenges, thus, contributing to gain a more accurate, high-throughput, and almost autonomous view of cellular structure and functions.
The proposed algorithm is able to identify and segment the structure of cells using AHE image reconstruction, image filtration, data augmentation, and improved watershed segmentation. Such a fusion of processing from different input images allows retrieving the structure of the required cells automatically.
